ProteinMPNN Execution Guide
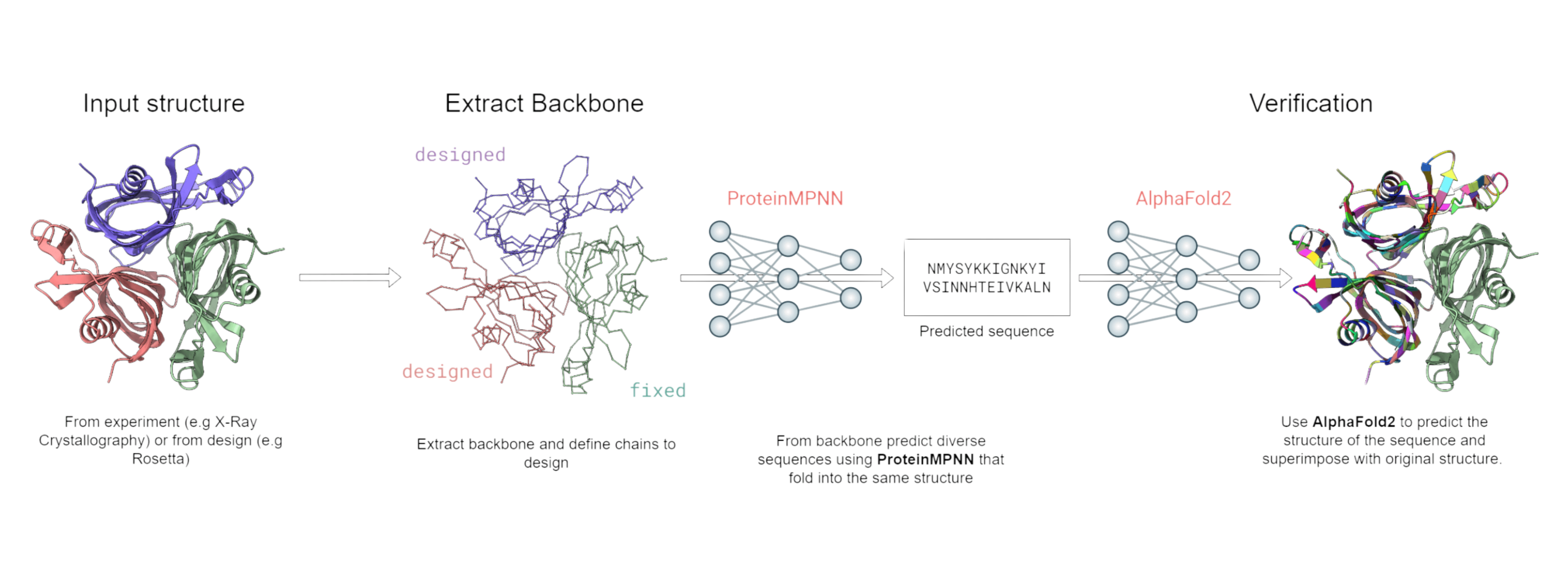
ProteinMPNN is a tool for protein sequence design based on backbone structures. Follow these steps on an E2E GPU Node.
Step 1: Clone Repository
git clone https://github.com/dauparas/ProteinMPNN.git
cd ProteinMPNN
Step 2: Create Environment
conda create --name mlfold
source activate mlfold
Step 3: Install Dependencies
For CPU:
pip install torch torchvision torchaudio --index-url https://download.pytorch.org/whl/cpu
For GPU:
pip install torch torchvision torchaudio --index-url https://download.pytorch.org/whl/cu121
conda install cudatoolkit=11.3
Step 4: Load Model
Update paths and load the pretrained weights:
path_to_model_weights=/mnt/model/vdb/ProteinMPNN/vanilla_model_weights
model_name="v_48_020"
checkpoint_path=$path_to_model_weights/$model_name.pt
Example in Python:
import torch
from protein_mpnn_utils import ProteinMPNN
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
checkpoint = torch.load(checkpoint_path, map_location=device)
model = ProteinMPNN(num_letters=21, hidden_dim=128, num_encoder_layers=3, num_decoder_layers=3)
model.load_state_dict(checkpoint['model_state_dict'])
model.to(device).eval()
print("Model loaded")
Step 5: Run Design
Provide a PDB file and specify designed/fixed chains.
Example:
pdb_path = "/mnt/model/vdb/ProteinMPNN/inputs/PDB_monomers/pdbs/5L33.pdb"
designed_chain = "A"
fixed_chain = ""
Adjust design options such as:
num_seqs = 1
sampling_temp = 0.1
batch_size = 1
Run the script to generate designed sequences.
Step 6: Output Results
The tool outputs designed sequences in FASTA format with scores and sequence recovery rates.
Example output:
>5L33, score=1.6044, designed_chains=['A'], model_name=v_48_020
HMPEEEKAARLFIEALEK...
Examples of PDBs
You can find Protein Data Bank (PDB) files at the following sources:
Use Cases
Expanding AlphaFold's Reach
AlphaFold is revolutionizing structural biology by unlocking previously elusive proteins. This breakthrough has significant applications in multiple fields:
Advancing Medicine
- Scientists are leveraging AlphaFold to develop novel treatments for diseases such as cancer, Alzheimer's, and antibiotic resistance.
- AI-driven protein structure predictions accelerate drug discovery and precision medicine.
Fighting Plastic Pollution
- Researchers are engineering enzymes that efficiently break down plastic waste, offering a potential solution to environmental crises.
- These advancements could lead to a sustainable approach to plastic degradation and waste management.
Improving Global Food Security
- Mapping plant proteins helps in developing drought-resistant crops.
- AI-driven insights contribute to more sustainable agricultural practices, enhancing global food supply.
AI-Driven Life Sciences
Machine learning is being integrated with physics-based simulations to push the boundaries of biological research:
- Curing diseases in months, not years – Accelerating the development of life-saving treatments.
- Beating superbugs before they strike – Predicting antibiotic resistance mechanisms to stay ahead of evolving pathogens.
- Engineering enzymes that devour plastic waste – Transforming waste management through bioengineering solutions.
AlphaFold's potential in structural biology, drug discovery, and AI-driven life sciences is reshaping how we approach global challenges, making groundbreaking scientific advancements accessible in record time.